K-State Researchers Find New Clues for Improving Wheat
Originally published by K-State Research and Extension;
Reprinted with permission from the Kansas Wheat Commission
A team of Kansas State University wheat scientists are tapping into 10,000 years of evolution in the plant’s genetic code as part of their continued efforts to understand how historic processes that shaped modern wheat can help to improve the varieties grown by today’s farmers.
The exhaustive study, which is published in Nature Genetics, involved sequencing the genomes of nearly 1,000 wheat lines collected from different parts of the world with different environments. The work was led by researchers from K-State and Agriculture Victoria of Australia, in collaboration with the University of Saskatchewan (Canada) and the University of Minnesota.
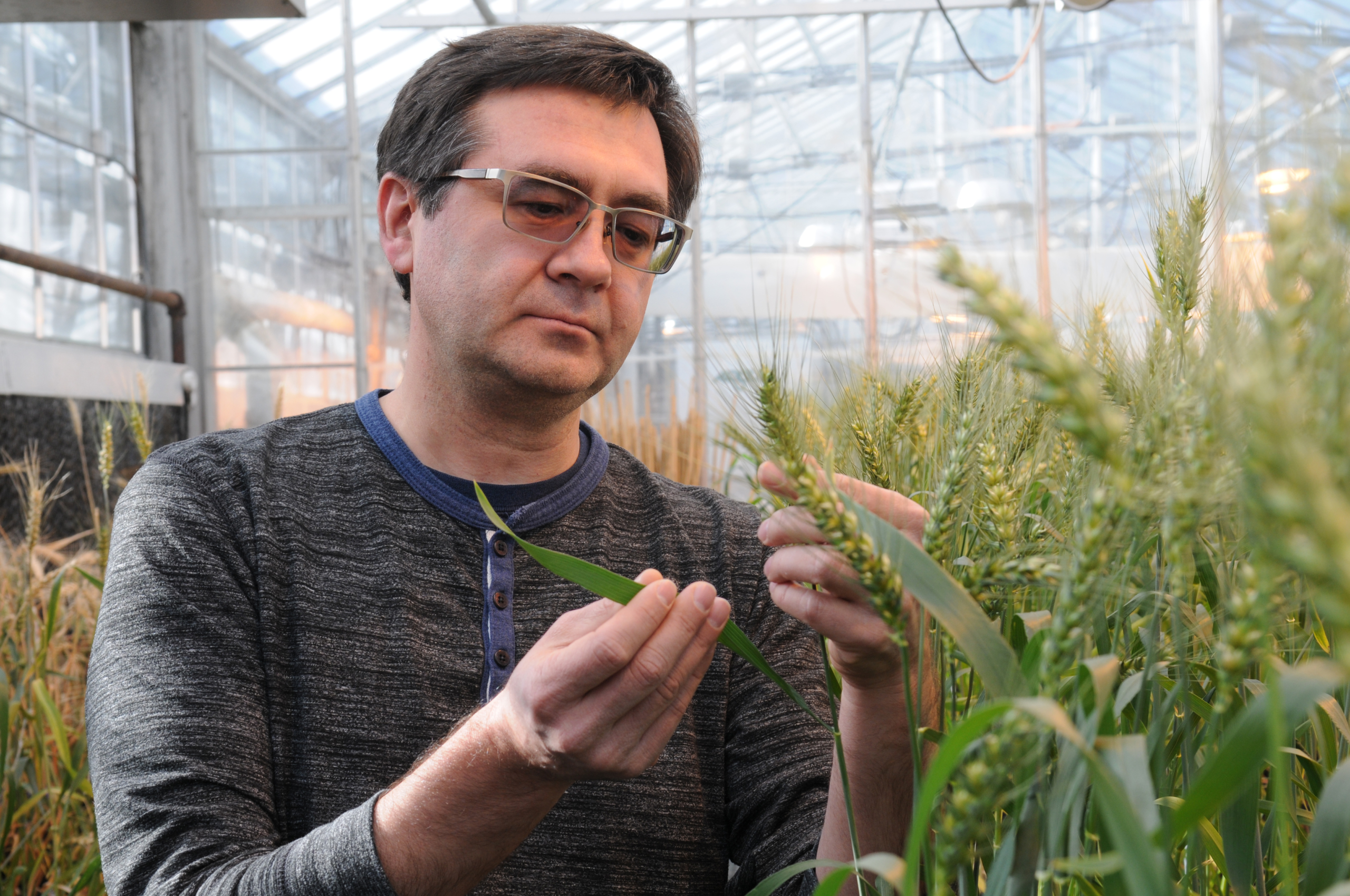
“We compared the genomes (in the 1,000 wheat lines) against each other, and looked for nucleotide base changes, or mutations, that distinguish one wheat accession from another,” said Eduard Akhunov, a K-State wheat geneticist.
He noted that the researchers found more than 7 million differences in the genetic code of the 1,000 lines.
“These differences can affect the function of genes that control various traits in wheat that helped it adapt to new growth conditions, such as withstanding drought and heat stresses; fighting off diseases; and yielding nutritious grain,” Akhunov said.
The changes that occurred in the genetic code can tell researchers a history of each wheat accession.
“When humans started spreading wheat from the site of its origin to other places, they brought it into contact with wild wheat, and wild ancestors accidentally began to inter-breed with bread wheat,” Akhunov said. “What happened then was that bread wheat inherited the genetic diversity that was present in the wild emmer wheat.”
That process of one species sharing genes with another species is called gene flow, and it is key for explaining the genetic diversity of today’s wheat varieties, according to K-State wheat breeder Allan Fritz.
“Understanding gene flow between wild emmer and common wheat is more than just academically interesting,” Fritz said. “The importance of historical introgression suggests that a more strategic use of wild emmer should have value for future wheat improvement.”
Fritz noted that K-State scientists have been using wild emmer in developing germplasm for new wheat varieties in projects funded by the Kansas Wheat Commission and the university’s Wheat Genetics Resource Center.
The work by Akhunov and his research team allows breeders to “evaluate the diversity in wild emmer and be intentional and strategic” in how they employ desired traits in new wheat varieties, according to Fritz.
“As we move forward, we can apply what has been learned here to also focus future efforts on traits related to health and nutrition that wouldn’t have been direct targets of historical selection,” he said.
Akhunov adds: “For the first time, we have described how wild emmer’s genetic diversity contributed to the development of bread wheat. And what it’s done since humans domesticated wheat is it’s helped to develop a better crop.”
K-State’s study was funded by the Agriculture and Food Research Initiative’s competitive grants program, administered through the U.S. Department of Agriculture’s National Institute of Food and Agriculture and part of the International Wheat Yield Partnership, which Akhunov said aims at increasing the genetic yield potential of wheat using innovative approaches.
Akhunov also said that Corteva Agriscience and the agriculture division of Dow/DuPont provided financial support through its collaboration with Agriculture Victoria Service. Their support, he said, allowed the researchers access to needed technologies and to develop the set of data indicating the genetic differences in wheat varieties, also called an SNP dataset.
K-State received additional funding from the Kansas Wheat Commission and the Bill and Melinda Gates Foundation.
Header Photo Caption: Kansas State University wheat geneticist and pathologist Eduard Akhunov works in the university’s greenhouse. Photo Credit: K-State Research & Extension